X-ray structure and mechanism of RNA polymerase II stalled at an antineoplastic monofunctional platinum-DNA adduct | PNAS
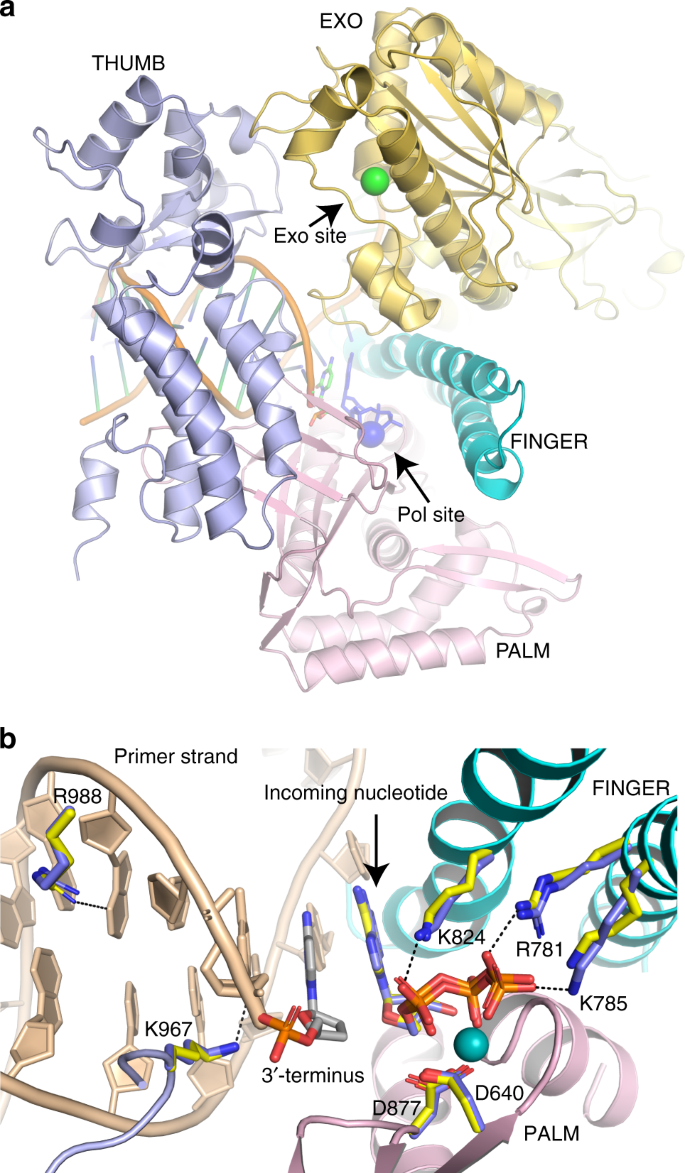
Structural consequence of the most frequently recurring cancer-associated substitution in DNA polymerase ε | Nature Communications

Diffusion of nucleoside triphosphates and role of the entry site to the RNA polymerase II active center | PNAS

POL partners with Army fuels specialists in aircraft refueling initiative > U.S. Air Forces Central > Article Display
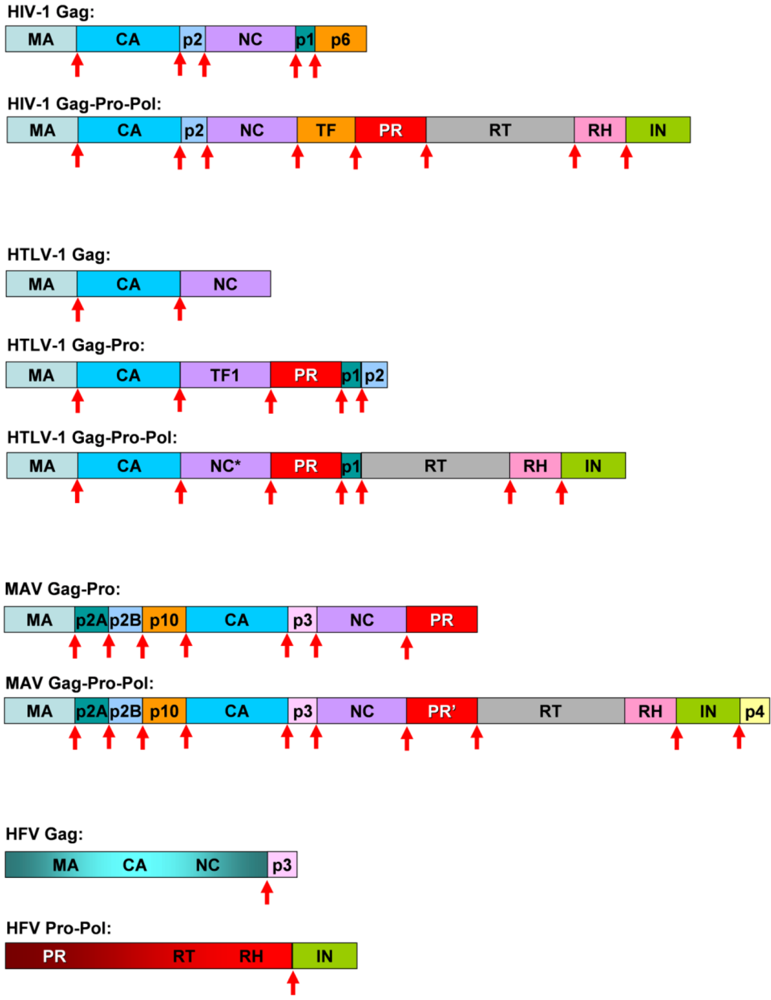
Viruses | Free Full-Text | Comparative Studies on Retroviral Proteases: Substrate Specificity | HTML

Integrator is recruited to promoter‐proximally paused RNA Pol II to generate Caenorhabditis elegans piRNA precursors | The EMBO Journal
Mammalian Base Excision Repair: Functional Partnership between PARP-1 and APE1 in AP-Site Repair | PLOS ONE

Variations in nuclear localization strategies among pol X family enzymes - Kirby - 2018 - Traffic - Wiley Online Library

Genetic evidence for reconfiguration of DNA polymerase θ active site for error-free translesion synthesis in human cells - Journal of Biological Chemistry

Primary sequence and structural conservation of the Pol ␦ active site.... | Download Scientific Diagram

Coordination of RNA Polymerase II Pausing and 3′ End Processing Factor Recruitment with Alternative Polyadenylation | Molecular and Cellular Biology

Structure and function of the pol II transcription machinery | Taatjes Lab | University of Colorado Boulder
Metal A and Metal B Sites of Nuclear RNA Polymerases Pol IV and Pol V Are Required for siRNA-Dependent DNA Methylation and Gene Silencing | PLOS ONE

Alternative DNA secondary structure formation affects RNA polymerase II promoter-proximal pausing in human | Genome Biology | Full Text
Widespread Backtracking by RNA Pol II Is a Major Effector of Gene Activation, 5′ Pause Release, Termination, and Trans